Cell Image Analysis
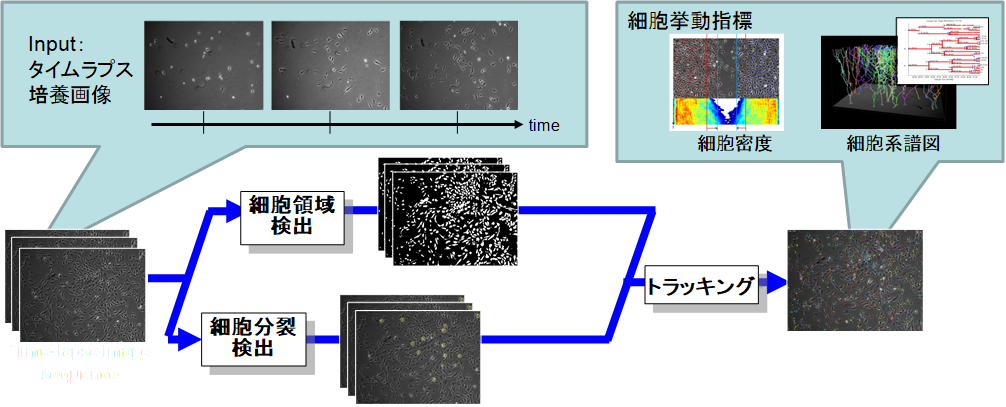
Advances in optics and imaging systems have enabled biologists to visualize a living specimen's dynamic pro-cesses by time-lapse microscopy images. However, the image data recorded during even a single experiment may consist of hundreds of objects over thousands of images, which makes manual inspection a tedious and inaccurate option. We addresses the need for computational tools and community resources that can enable the quantitative system-level biology automatically. The domain of our current interests are cell image analysis algorithms including microscopy image restoration, cell mitotic event detection and cell tracking by data association, which are integrated into an online automated system.
Cell Segmentation and Tracking
Cell Tracking with Deep Learning for Cell Detection and Motion Estimation in Low-Frame-Rate
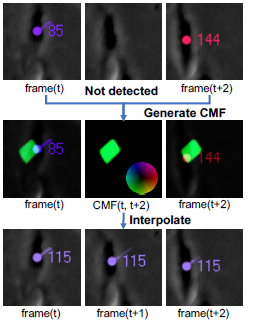
Cell behavior analysis in High-throughput biological experiments is important for research and discovery in biology and medicine. To perform the high-throughput experiments, it requires to capture images in low frame rate in order to recode images on multi-points. In such low frame rate image sequence, movements of cells between successive frames are often larger than distances to nearby cells, and thus current methods based on proximity do not work properly. In this study, we propose a cell tracking method that enables to track cells in low frame rate by simultaneously estimating all of the cell motions in an image. In the experiments under dense conditions in low frame rate, our method outperformed the other methods.
- [J. Hayashida and R. Bise, MICCAI,2019, accepted]
Weakly Supervised Cell Instance Segmentation in Dense by Propagating from Detection Map
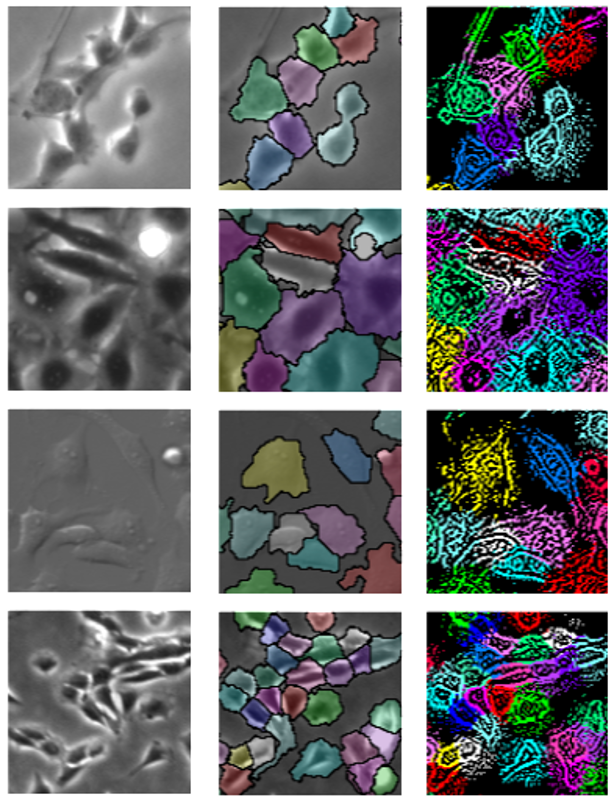
Cell shape analysis is important in biomedical research. Deep learning methods may perform to segment individual cells if they use suffcient training data that the boundary of each cell is annotated. However, it is very time-consuming for preparing such detailed annotation for many cell culture conditions. In this paper, we propose a weakly supervised method that can segment individual cell regions who touch each other with unclear boundaries in dense conditions without the training data for cell regions. We demonstrated the effecacy of our method using several data-set including multiple cell types captured by several types of microscopy. Our method achieved the highest accuracy compared with several conventional methods. In addition, we demonstrated that our method can perform without any annotation by using fluorescence images that cell nuclear were stained as training data.
- [K. Nishimura, E.D.F. Ker, and R. Bise, MICCAI,2019, accepted]
Cell Detection Method from Redundant Candidates under the Non-Overlapping Constraints
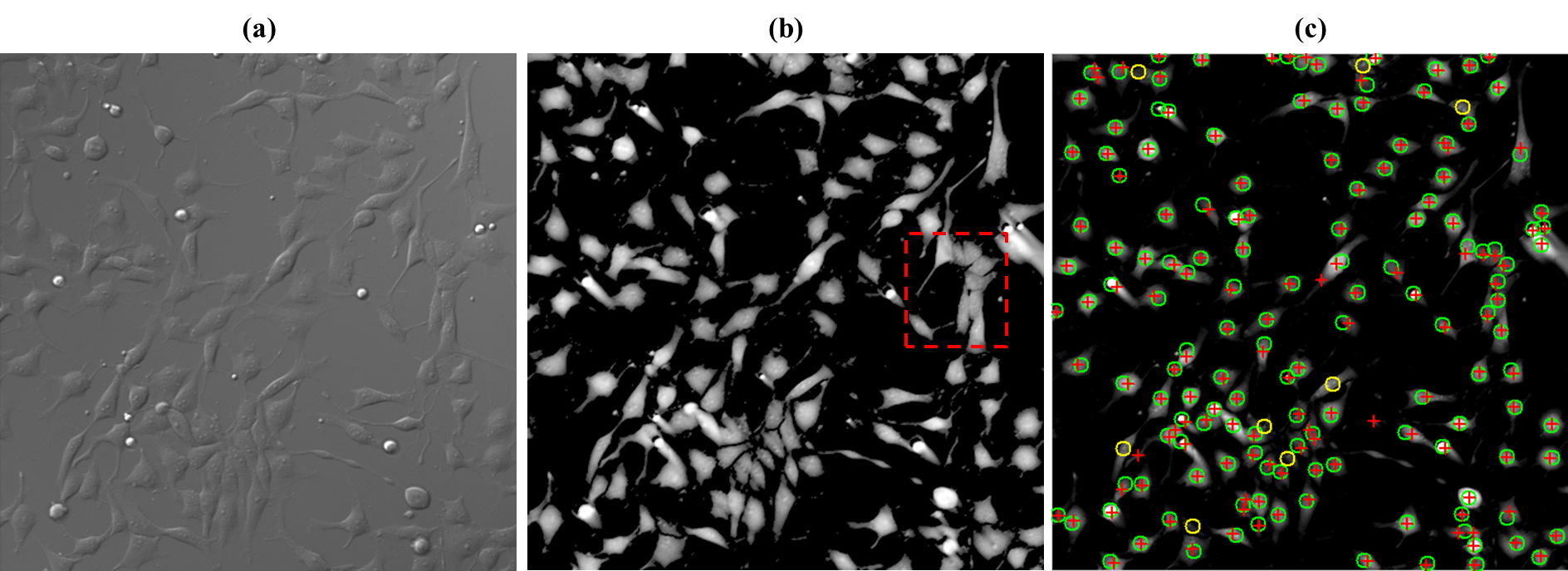
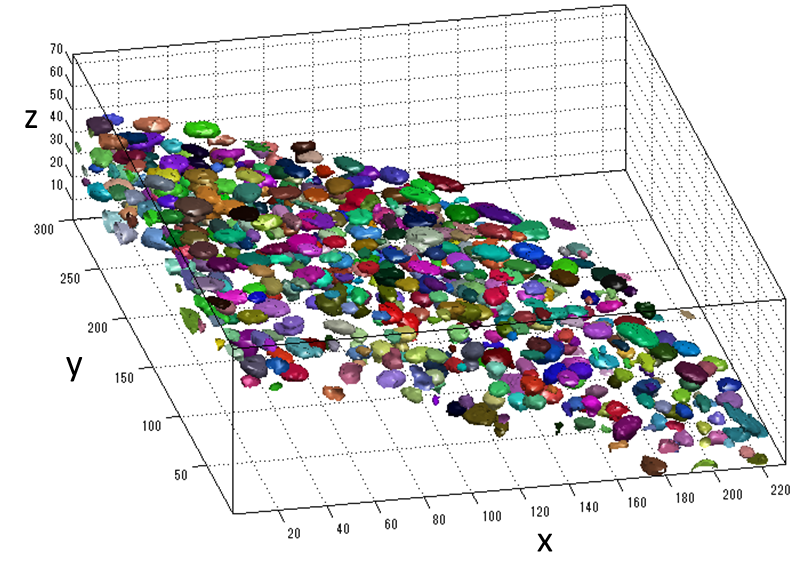
Cell detection in microscopy images is essential for automated cell behavior analysis including cell shape analysis and cell tracking. Robust cell detection in high-density and lowcontrast images is still challenging since cells often touch and partially overlap, forming a cell cluster with blurry intercellular boundaries. In such cases, current methods tend to detect multiple cells as a cluster. If the control parameters are adjusted to separate the touching cells, other problems often occur: a single cell may be segmented into several regions, and cells in lowintensity regions may not be detected. To solve these problems, we first detect redundant candidate regions, which include many false positives but in turn very few false negatives, by allowing candidate regions to overlap with each other. Next, the score for how likely the candidate region contains the main part of a single cell is computed for each cell candidate using supervised learning. Then we select an optimal set of cell regions from the redundant regions under non-overlapping constraints, where each selected region looks like a single cell and the selected regions do not overlap. We formulate this problem of optimal region selection as a binary linear programming problem under non-overlapping constraints. We demonstrated the effectiveness of our method for several types of cells in microscopy images. Our method performed better than five representative methods, achieving an F-measure of over 0.9 for all data sets. Experimental application of the proposed method to 3D images demonstrated that also works well for 3D cell detection.
Cell Segmentation in Microscopy Imagery Using a Bag of Local Bayesian Classifiers
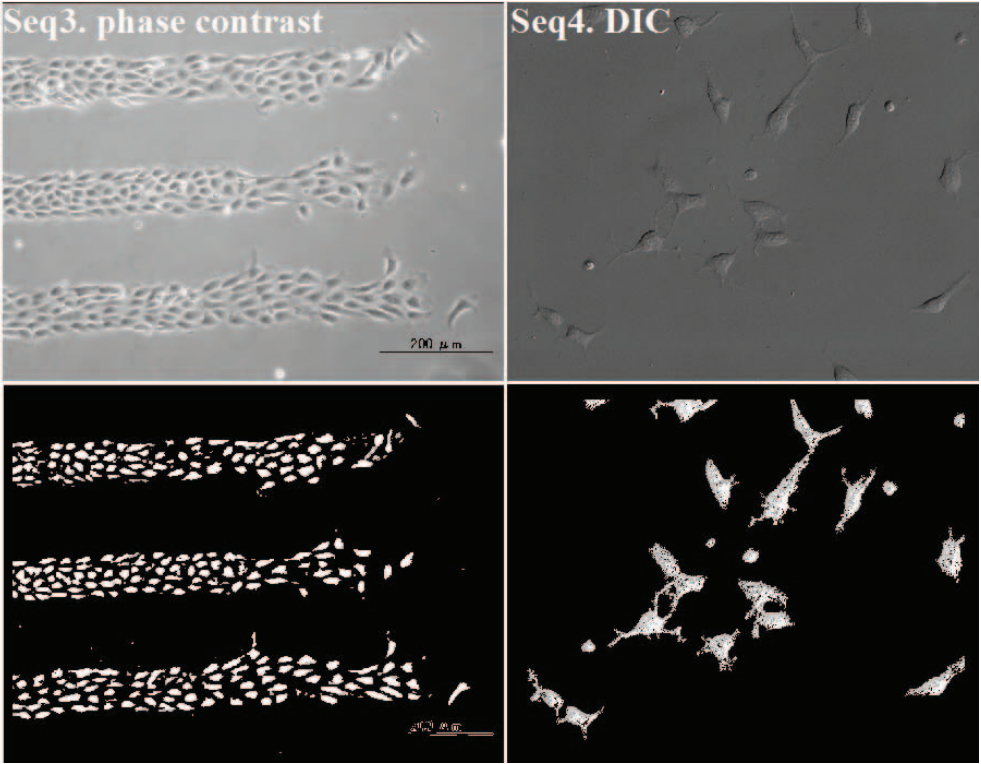
Cell segmentation in microscopy imagery is essential for many bioimage applications such as cell tracking. To segment cells from the background accurately, we present a pixel classification approach that is independent of cell type or imaging modality. We train a set of Bayesian classifiers from clustered local training image patches. Each Bayesian classifier is an expert to make decision in its specific domain. The decision from the mixture of experts determines how likely a new pixel is a cell pixel. We demonstrate the effectiveness of this approach on four cell types with diverse morphologies under different microscopy imaging modalities.
Detection of Hematopoietic Stem Cells in Microscopy Images Using a Bank of Ring Filters
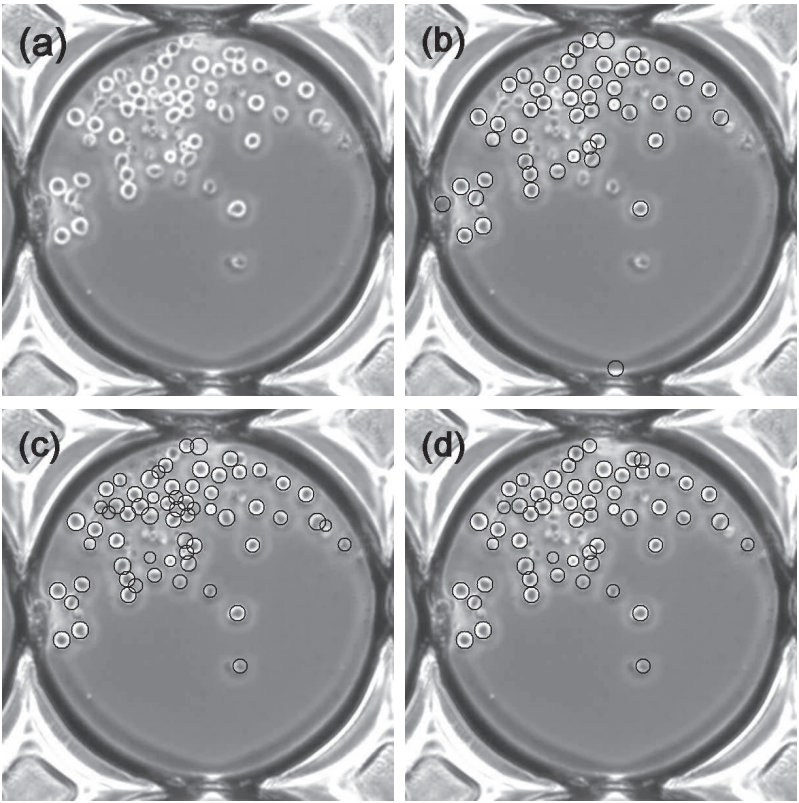
We present a method for robustly detecting hematopoietic stem cells (HSCs) in phase contrast microscopy images. HSCs appear to be easy to detect since they typically appear as round objects. However, when HSCs are touching and overlapping, showing the variations in shape and appearance, standard pattern detection methods, such as Hough transform and correlation, do not perform well. The proposed method exploits the output pattern of a ring filter bank applied to the input image, which consists of a series of matched filters with multiple-radius ring-shaped templates. By modeling the profile of each filter response as a quadratic surface, we explore the variations of peak curvatures and peak values of the filter responses when the ring radius varies. The method is validated on thousands of phase contrast microscopy images with different acquisition settings, achieving 96.5% precision and 94.4% recall.
Mitosis Detection
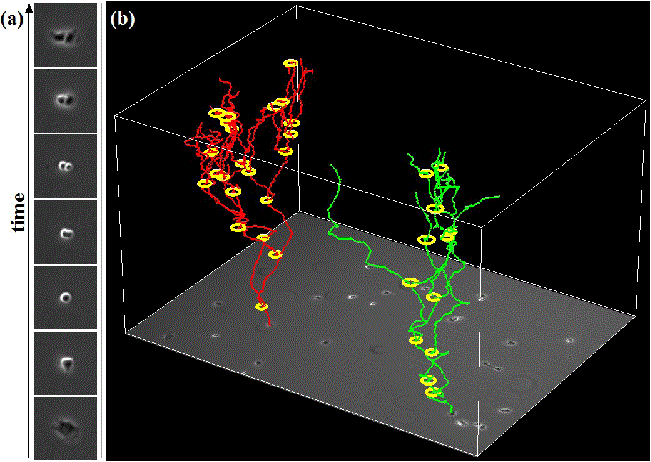
We propose an effective approach for automated mitosis detection using phase-contrast time-lapse microscopy, which is a non-destructive imaging modality, thereby allowing continuous monitoring of cells in culture. In our approach, we present a probabilistic model for event detection, which can simultaneously 1) identify spatio-temporal patch sequences that contain a mitotic event and 2) localize a birth event, defined as the time and location at which cell division is completed and two daughter cells are born. Our approach significantly outperforms previous approaches in terms of both detection accuracy and computational efficiency, when applied to multipotent C3H10T1/2 mesenchymal and C2C12 myoblastic stem cell populations.
Cell Tracking based on Frame by Frame Data Association
The image segmentation algorithm segments blobs from input images that can be individual cells or cell clusters (overlapped cells), and the mitosis detection algorithm locates birth events where and when one cell divides into two cells. Based on the outputs of these two algorithms, we developed a cell-blob correspondence algorithm performing data association between the cells in the previous frame and the blobs segmented in the current frame. In detail, the association algorithm makes the following hypotheses with corresponding likelihood for all possible cell actions.
Cell Tracking based on Global Data Association
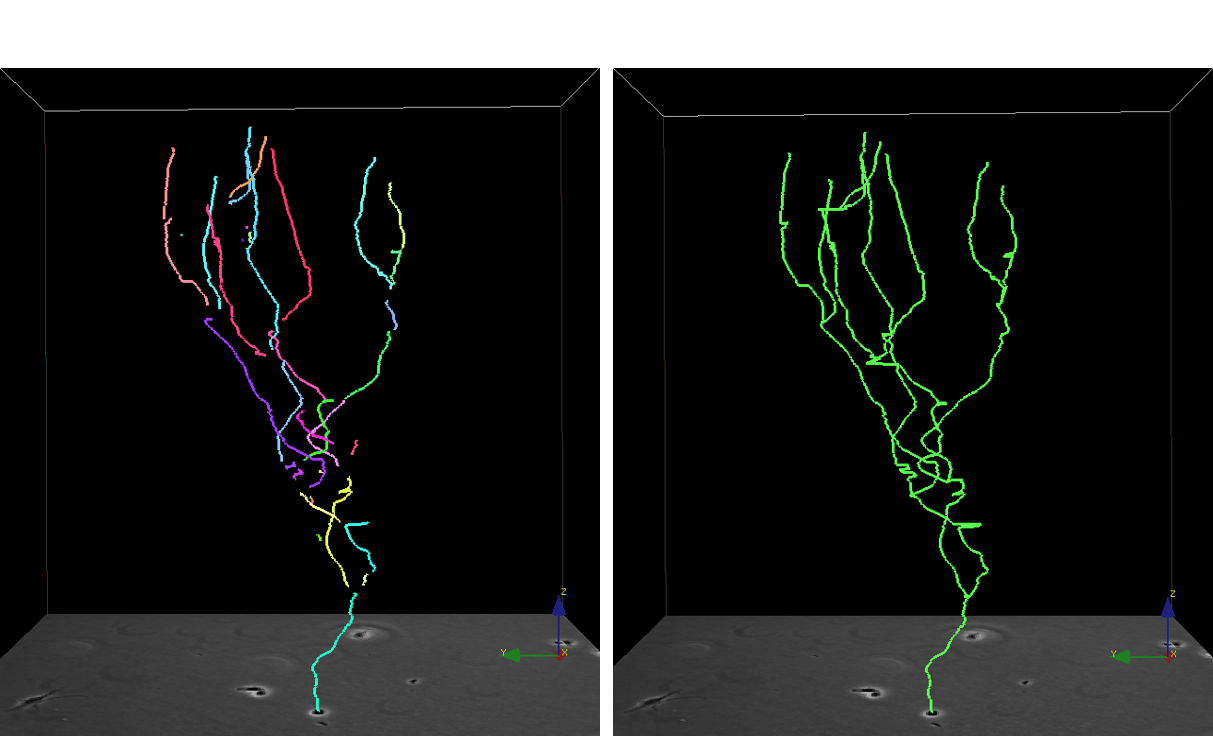

Automated cell tracking in populations is important for research and discovery in biology and medicine. We propose a cell tracking method based on global spatiotemporal data association which considers hypotheses of initialization, termination, translation, division and false positive in an integrated formulation. Firstly, reliable tracklets (i.e., short trajectories) are generated by linking detection responses based on frame-by-frame association. Next, these tracklets are globally associated over time to obtain final cell trajectories and lineage trees. During global association, tracklets form tree structures where a mother cell divides into two daughter cells. We formulate the global association for tree structures as a maximum-a-posteriori (MAP) problem and solve it by linear programming. This approach is quantitatively evaluated on sequences with thousands of cells captured over several days.
Application
Wound Healing Assay
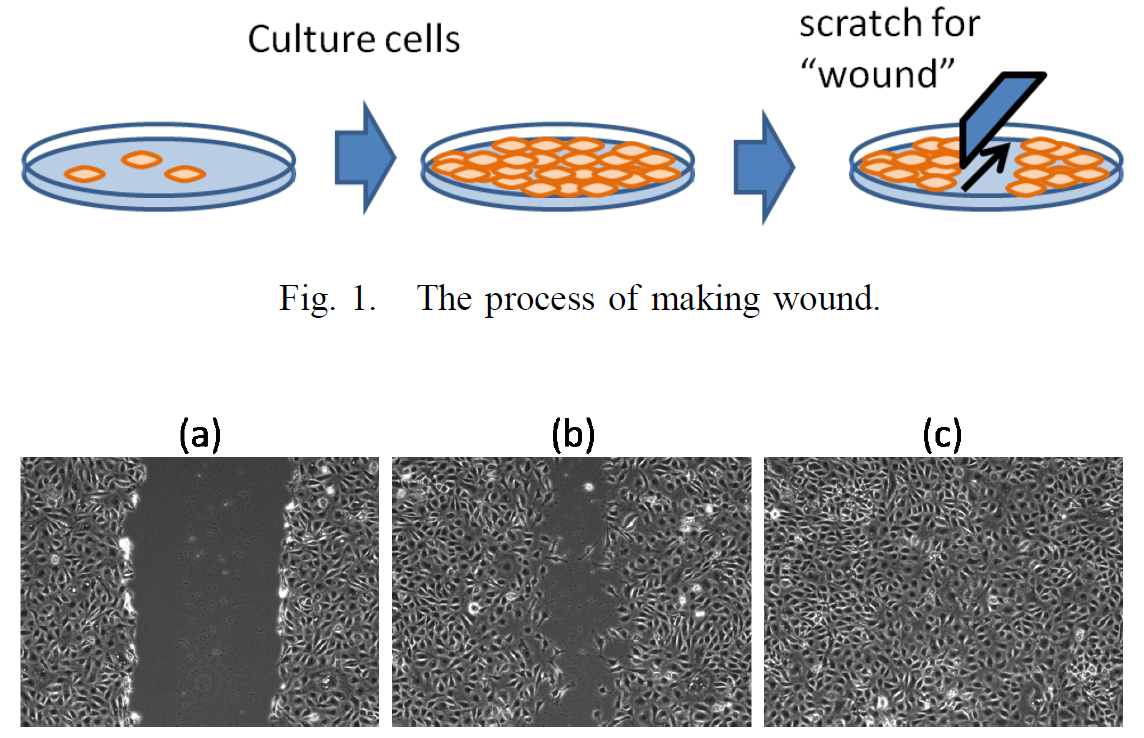
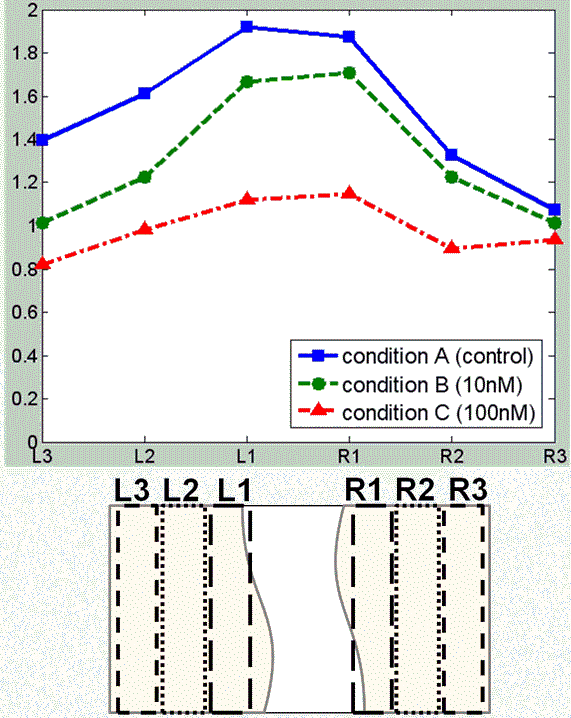
The wound healing assay in vitro is widely used for research and discovery in biology and medicine. This assay allows for observing the healing process in vitro in which the cells on the edges of the artificial wound migrate toward the wound area. The influence of different culture conditions can be measured by observing the change in the size of the wound area. For further investigation, more detailed measurements of the cell behaviors are required. We present an application of automatic cell tracking in phase-contrast microscopy images to wound healing assay. The cell behaviors under three different culture conditions have been analyzed. Our cell tracking system can track individual cells during the healing process and provide detailed spatio-temporal measurements of cell behaviors including cell density, cell migration speed and direction, and the statistics of cell mitosis events. The application demonstrates the effectiveness of automatic cell tracking for quantitative and detailed analysis of the cell behaviors in wound healing assay in vitro.
Tracing behavior of endothelial cells promotes vascular network formation
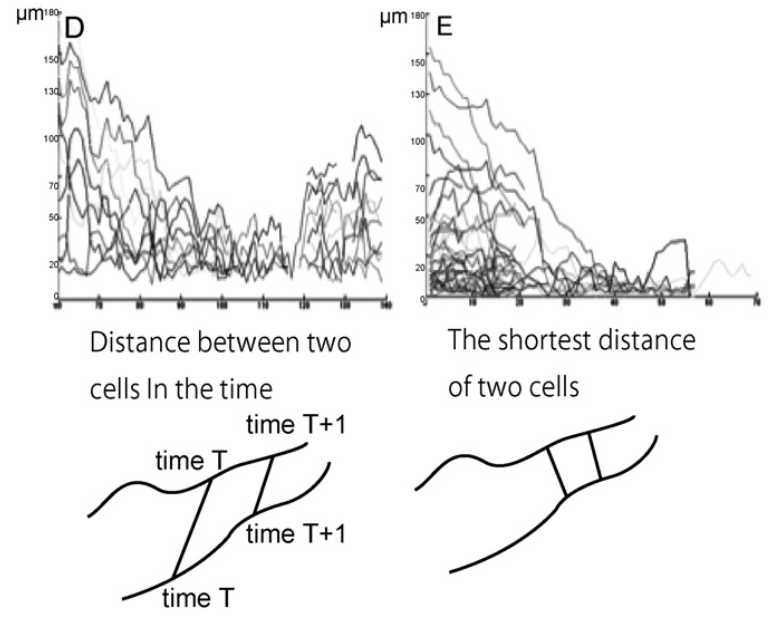
The in vitro formation of network structures derived from endothelial cells in grafts before transplantation contributes to earlier engraftment. In a previous study, endothelial cells migrated to form a net-shaped structure in co-culture. However, the specific network formation behavior of endothelial cells during migration remains unclear. In this study, we demonstrated the tracing behavior and cell cycle of endothelial cells using Fucci-labeled (Fluorescent Ubiquitination-based Cell Cycle Indicator) endothelial cells. Here, we observed the co-culture of Fucci-labeled human umbilical vein endothelial cells (HUVECs) together with normal human dermal fibroblasts (NHDFs) using time-lapse imaging and analyzed by multicellular concurrent tracking. In the G0/G1 period, HUVECs migrate faster than in the S/G2/M period, because G0/G1 is the mobile phase and S/G2/M is the proliferation phase in the cell cycle. When HUVECs are co-cultured, they tend to move randomly until they find existing tracks that they then follow to form clusters. Extracellular matrix (ECM) staining showed that collagen IV, laminin and thrombospondin deposited in accordance with endothelial cell networks. Therefore the HUVECs may migrate on the secreted ECM and exhibit tracing behavior, where the HUVECs migrate toward each other. These results suggested that ECM and a cell phase contributed to form a network by accelerating cell migration.
An engineered approach to stem cell culture: automating the decision process for real-time adaptive subculture of stem cells
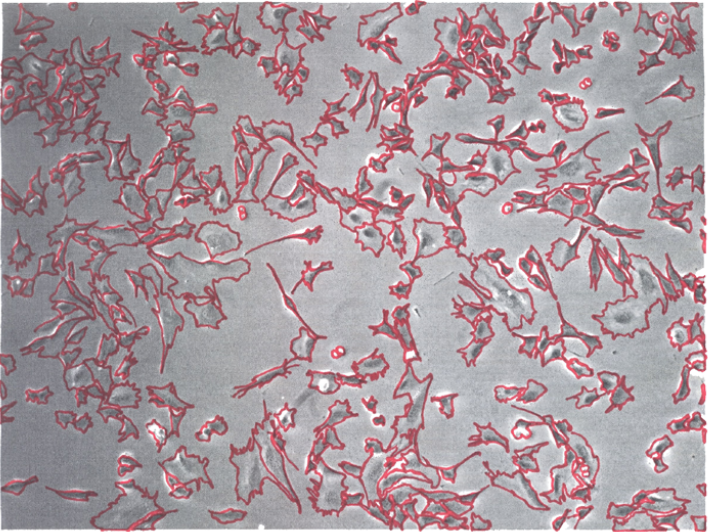
Current cell culture practices are dependent upon human operators and remain laborious and highly subjective, resulting in large variations and inconsistent outcomes, especially when using visual assessments of cell confluency to determine the appropriate time to subculture cells. Although efforts to automate cell culture with robotic systems are underway, the majority of such systems still require human intervention to determine when to subculture. Thus, it is necessary to accurately and objectively determine the appropriate time for cell passaging. Optimal stem cell culturing that maintains cell pluripotency while maximizing cell yields will be especially important for efficient, cost-effective stem cell-based therapies. Toward this goal we developed a real-time computer vision-based system that monitors the degree of cell confluency with a precision of 0.791±0.031 and recall of 0.559±0.043. The system consists of an automated phase-contrast time-lapse microscope and a server. Multiple dishes are sequentially imaged and the data is uploaded to the server that performs computer vision processing, predicts when cells will exceed a pre-defined threshold for optimal cell confluency, and provides a Web-based interface for remote cell culture monitoring. Human operators are also notified via text messaging and e-mail 4 hours prior to reaching this threshold and immediately upon reaching this threshold. This system was successfully used to direct the expansion of a paradigm stem cell population, C2C12 cells. Computer-directed and human-directed control subcultures required 3 serial cultures to achieve the theoretical target cell yield of 50 million C2C12 cells and showed no difference for myogenic and osteogenic differentiation. This automated vision-based system has potential as a tool toward adaptive real-time control of subculturing, cell culture optimization and quality assurance/quality control, and it could be integrated with current and developing robotic cell cultures systems to achieve complete automation.
Mechanical characterization of adult stem cells from bone marrow and perivascular niches
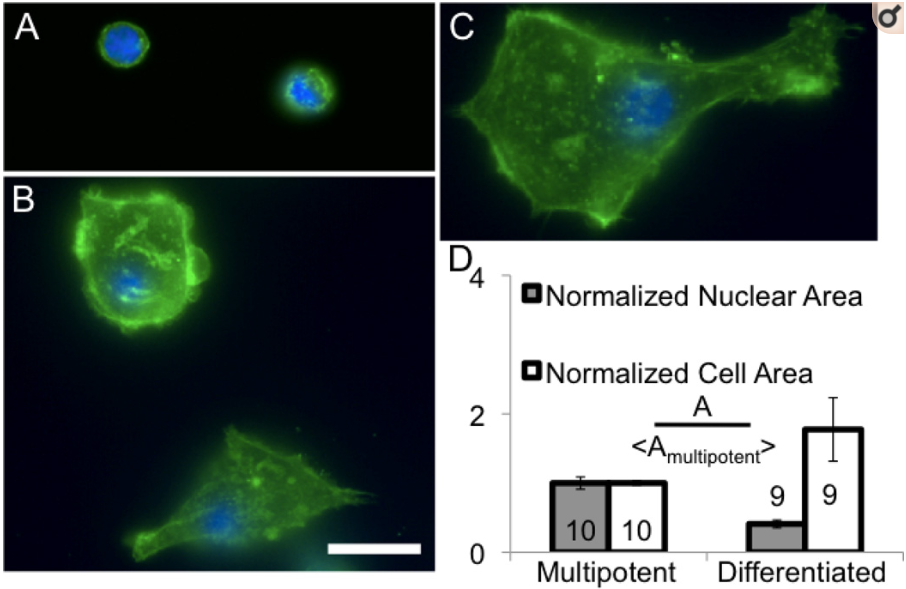
Therapies using adult stem cells often require mechanical manipulation such as injection or incorporation into scaffolds. However, force-induced rupture and mechanosensitivity of cells during manipulation is largely ignored. Here, we image cell mechanical structures and perform a biophysical characterization of three different types of human adult stem cells: bone marrow CD34+ hematopoietic, bone marrow mesenchymal and perivascular mesenchymal stem cells. We use micropipette aspiration to characterize cell mechanics and quantify deformation of subcellular structures under force and its contribution to global cell deformation. Our results suggest that CD34+ cells are mechanically suitable for injection systems since cells transition from solid- to fluid-like at constant aspiration pressure, probably due to a poorly developed actin cytoskeleton. Conversely, mesenchymal stem cells from the bone marrow and perivascular niches are more suitable for seeding into biomaterial scaffolds since they are mechanically robust and have developed cytoskeletal structures that may allow cellular stable attachment and motility through solid porous environments. Among these, perivascular stem cells cultured in 6% oxygen show a developed cytoskeleton but a more compliant nucleus, which can facilitate the penetration into pores of tissues or scaffolds. We confirm the relevance of our measurements using cell motility and migration assays and measure survival of injected cells. Since different types of adult stem cells can be used for similar applications, we suggest considering mechanical properties of stem cells to match optimal mechanical characteristics of therapies.
Data Set
Phase Contrast Time-Lapse Microscopy Datasets with Automated and Manual Cell Tracking Annotations
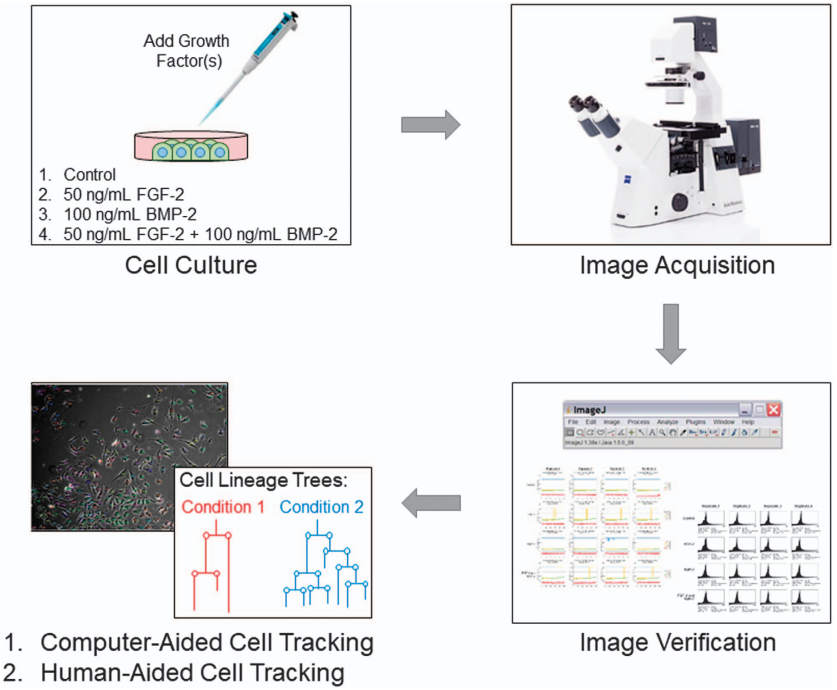
Phase contrast time-lapse microscopy is a non-destructive technique that generates large volumes of image-based information to quantify the behaviour of individual cells or cell populations. To guide the development of algorithms for computer-aided cell tracking and analysis, 48 time-lapse image sequences, each spanning approximately 3.5 days, were generated with accompanying ground truths for C2C12 myoblast cells cultured under 4 different media conditions, including with fibroblast growth factor 2 (FGF2), bone morphogenetic protein 2 (BMP2), FGF2?+?BMP2, and control (no growth factor). The ground truths generated contain information for tracking at least 3 parent cells and their descendants within these datasets and were validated using a two-tier system of manual curation. This comprehensive, validated dataset will be useful in advancing the development of computer-aided cell tracking algorithms and function as a benchmark, providing an invaluable opportunity to deepen our understanding of individual and population-based cell dynamics for biomedical research.
